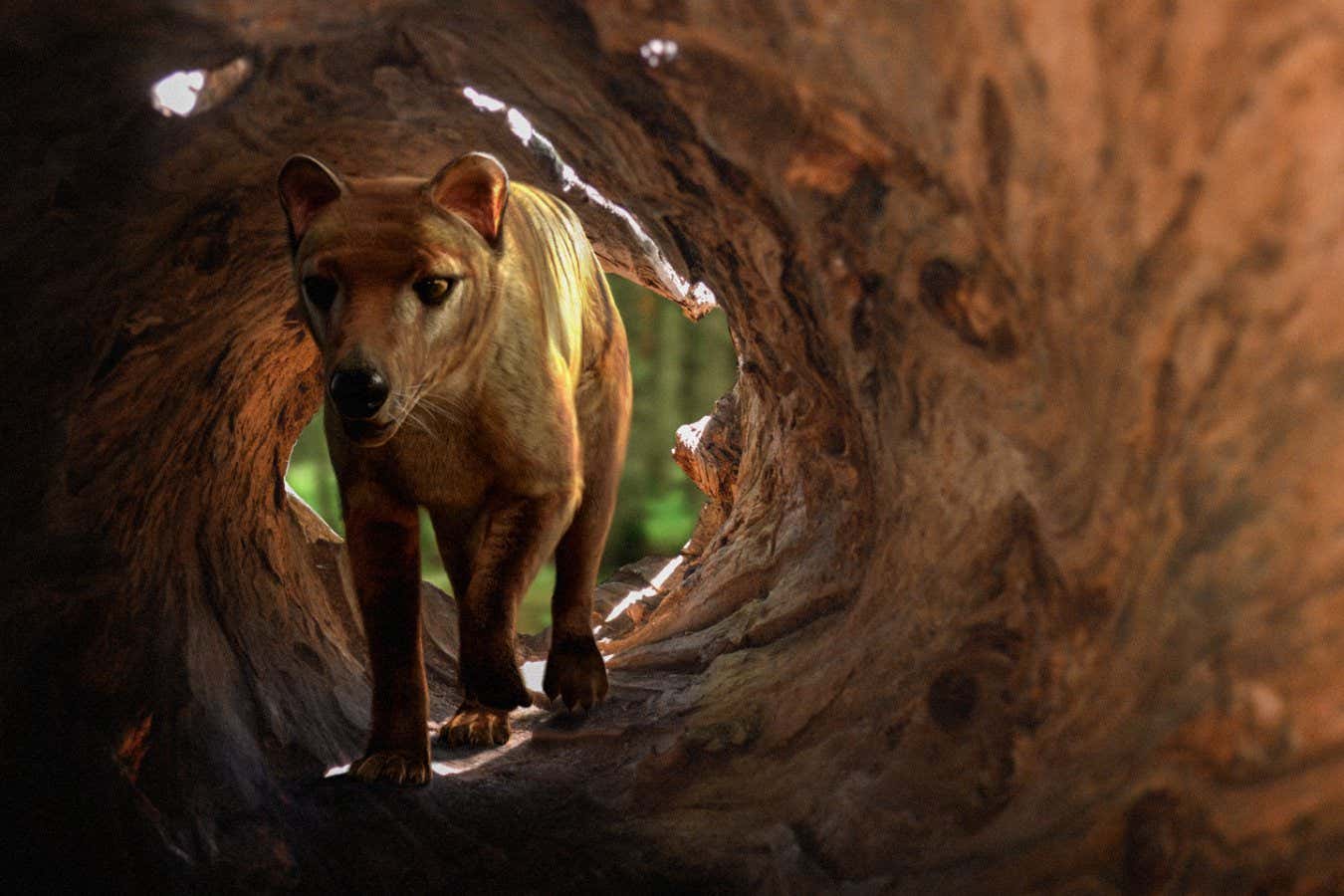
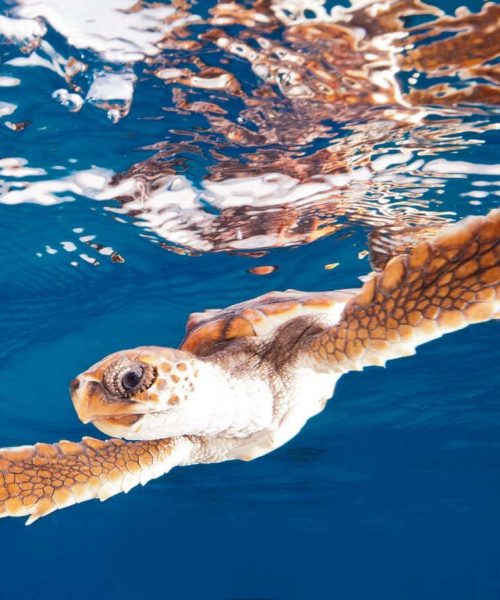
Thylacines, or Tasmanian tigers, went extinct in 1936
Colossal Biosciences
The genome of the extinct thylacine has been nearly completely sequenced, de-extinction company Colossal has announced. It says the genome is more than 99.9 per cent complete, with just 45 gaps that will soon be closed – but it has provided no evidence to back up its claim.
“It’s a fairly difficult thing to get a fully complete genome of almost any organism,” says Emilio Mármol-Sánchez at the University of Copenhagen, Denmark, whose team was the first to extract RNA from a preserved thylacine. For example, the last few holdouts of the human genome were only fully sequenced in the past few years.
Advertisement
Thylacines, also known as Tasmanian tigers, were carnivorous marsupials once found throughout Australia, but by the time European explorers arrived, they were limited to Tasmania. The last known thylacine died in a zoo in 1936.
The genome of a preserved thylacine was first sequenced in 2017 using tissue from a then-108-year-old thylacine pouch preserved in alcohol. However, this genome was far from complete, with many gaps. Now Colossal, which also aims to recreate the woolly mammoth, says it has largely completed this genome with the help of additional DNA from a 120-year-old tooth.
“Our genome is not as complete as the most complete human genome, but we were able to take advantage of some of the same technologies,” says Andrew Pask at the University of Melbourne in Australia, a member of Colossal’s scientific advisory board.
It is difficult to completely sequence the genomes of plants and animals because there are large sections where the same sequences are repeated many times. Standard techniques that sequence small segments of DNA at a time don’t work for these parts – it is like trying to reassemble a book from a list of the words in it.
Newer, long-read techniques can sequence much larger segments of DNA – whole pages of the book. However, old DNA usually breaks up into lots of small pieces, so these methods don’t often help.
“Most ancient samples preserve DNA fragments that are on the order of tens of bases long – hundreds if we are lucky,” says Pask. “The sample we were able to access was so well preserved that we could recover fragments of DNA that were thousands of bases long.”
Given the lack of any other thylacine genomes to make a comparison with, there is no direct way to tell how complete it is – instead Pask says Colossal is using other related species in the same family to make this estimate.
But even if the genome is as complete as Colossal thinks and it really can fill in the remaining gaps, there is currently no feasible way to generate living cells containing this genome. Instead, Colossal plans to genetically modify a living marsupial called the fat-tailed dunnart to make it more like a thylacine.
“It’s more a recreation of some traits,” says Mármol-Sánchez. “It would not be an extinct animal, but a pretty weird, modified version of the modern animal that resembles our image of those extinct animals.”
Colossal says it has made a record 300 genetic edits to the genomes of dunnart cells growing in culture. So far, all are small changes, but Pask says the team plans to swap in tens of thousands of base pairs of thylacine DNA in the near future. It isn’t yet clear how many edits will be required to achieve the company’s goal of recreating the thylacine, he says.
When asked why Colossal had provided no evidence in support of its claims, CEO Ben Lamm said the company’s sole focus is de-extinction, not writing scientific papers. “We are not an academic lab where papers are their main focus,” said Lamm. “We will continue to make progress much faster than the process of writing scientific papers.”
Topics: